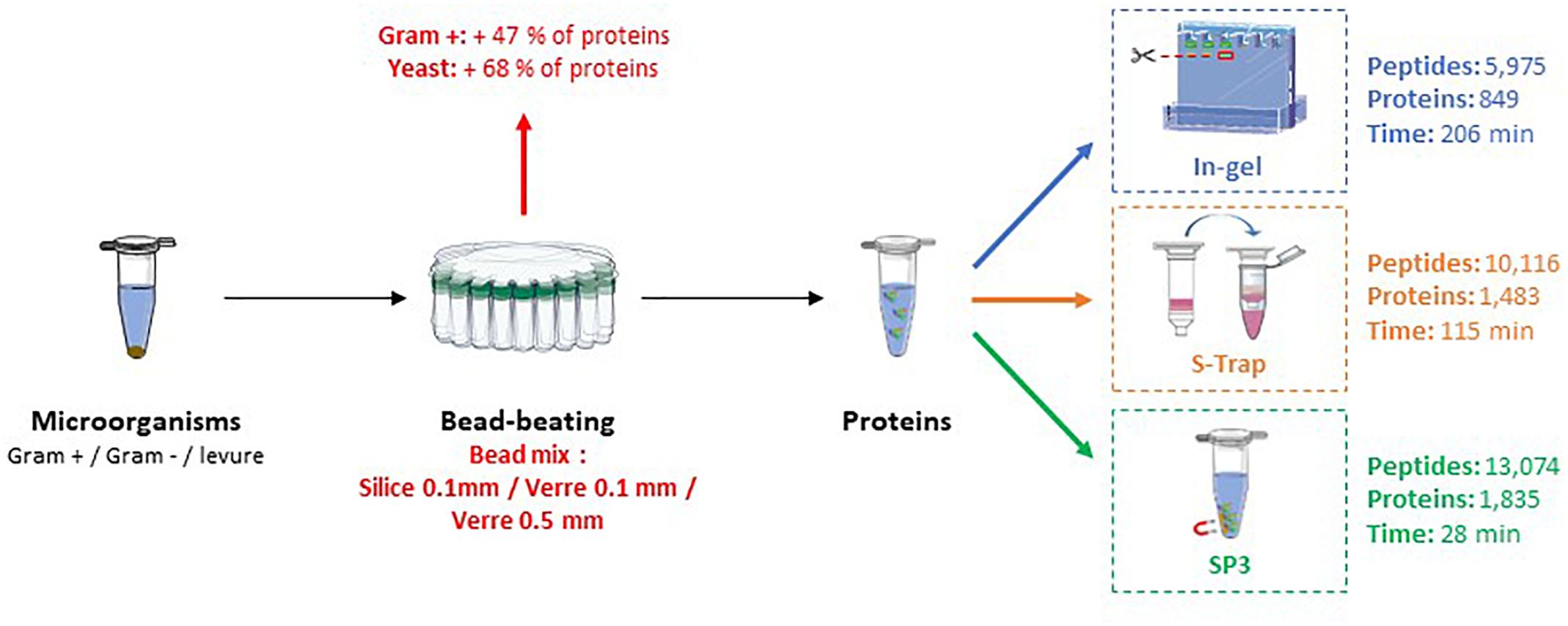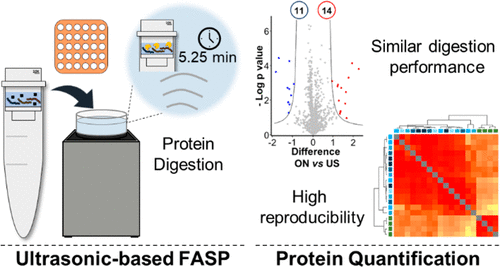
Ultrasonic-Based Filter Aided Sample Preparation as the General Method to Sample Preparation in Proteomics.,Analytical Chemistry - X-MOL
Overview of the analytical workflow: (A) Isolation of extracellular... | Download Scientific Diagram

Gel‐aided sample preparation (GASP)—A simplified method for gel‐assisted proteomic sample generation from protein extracts and intact cells - Fischer - 2015 - PROTEOMICS - Wiley Online Library

Precision Mapping of an In Vivo N-Glycoproteome Reveals Rigid Topological and Sequence Constraints: Cell

Reliable FASP-based procedures for optimal quantitative proteomic and phosphoproteomic analysis on samples from acute myeloid leukemia patients | Biological Procedures Online | Full Text
![PDF] Filter-Aided Sample Preparation: The Versatile and Efficient Method for Proteomic Analysis. | Semantic Scholar PDF] Filter-Aided Sample Preparation: The Versatile and Efficient Method for Proteomic Analysis. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/892cadcebd383567fff4f15e61571818b0ac1bdc/3-Figure1-1.png)
PDF] Filter-Aided Sample Preparation: The Versatile and Efficient Method for Proteomic Analysis. | Semantic Scholar

imFASP: An integrated approach combining in-situ filter-aided sample pretreatment with microwave-assisted protein digestion for fast and efficient proteome sample preparation - ScienceDirect

Enhanced filter-aided sample preparation (FASP) workflow. Samples are... | Download Scientific Diagram

Sample Preparation by Easy Extraction and Digestion (SPEED) - A Universal, Rapid, and Detergent-free Protocol for Proteomics based on Acid Extraction | bioRxiv
The Amicon Ultra-0.5 mL Centrifugal Filters used in the FASP procedure.... | Download Scientific Diagram
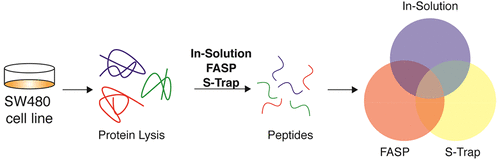
Mass Spec Master: In-solution, In-gel, FASP and S-Traping - a voyage through protein sample preparation
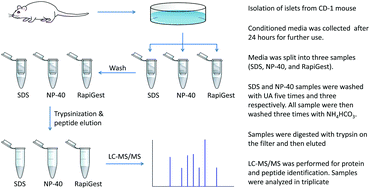
Sample preparation protocol for bottom-up proteomic analysis of the secretome of the islets of Langerhans - Analyst (RSC Publishing)

Enhanced FASP (eFASP) to increase proteome coverage and sample recovery for quantitative proteomic experiments. - Abstract - Europe PMC